
Visualization of structures
We offer following software dealing with protein structures visualization:
Mol*
Mol*VS
OverProt
2DProts
Mol*
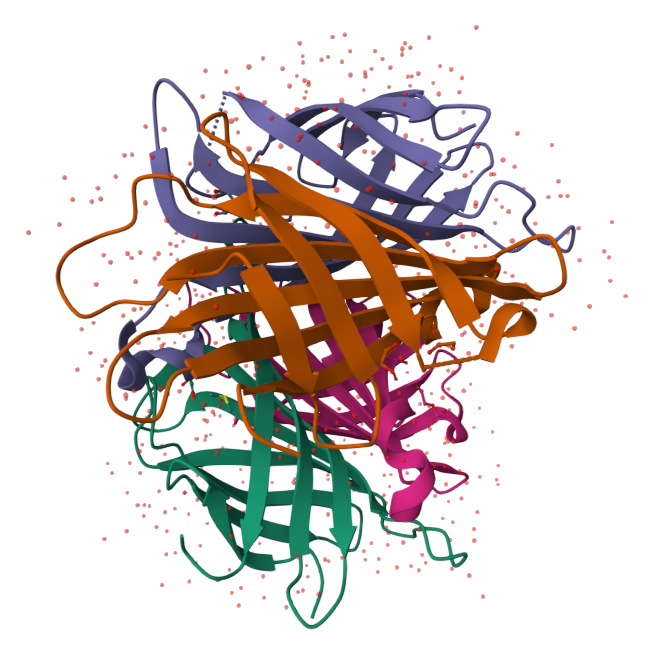
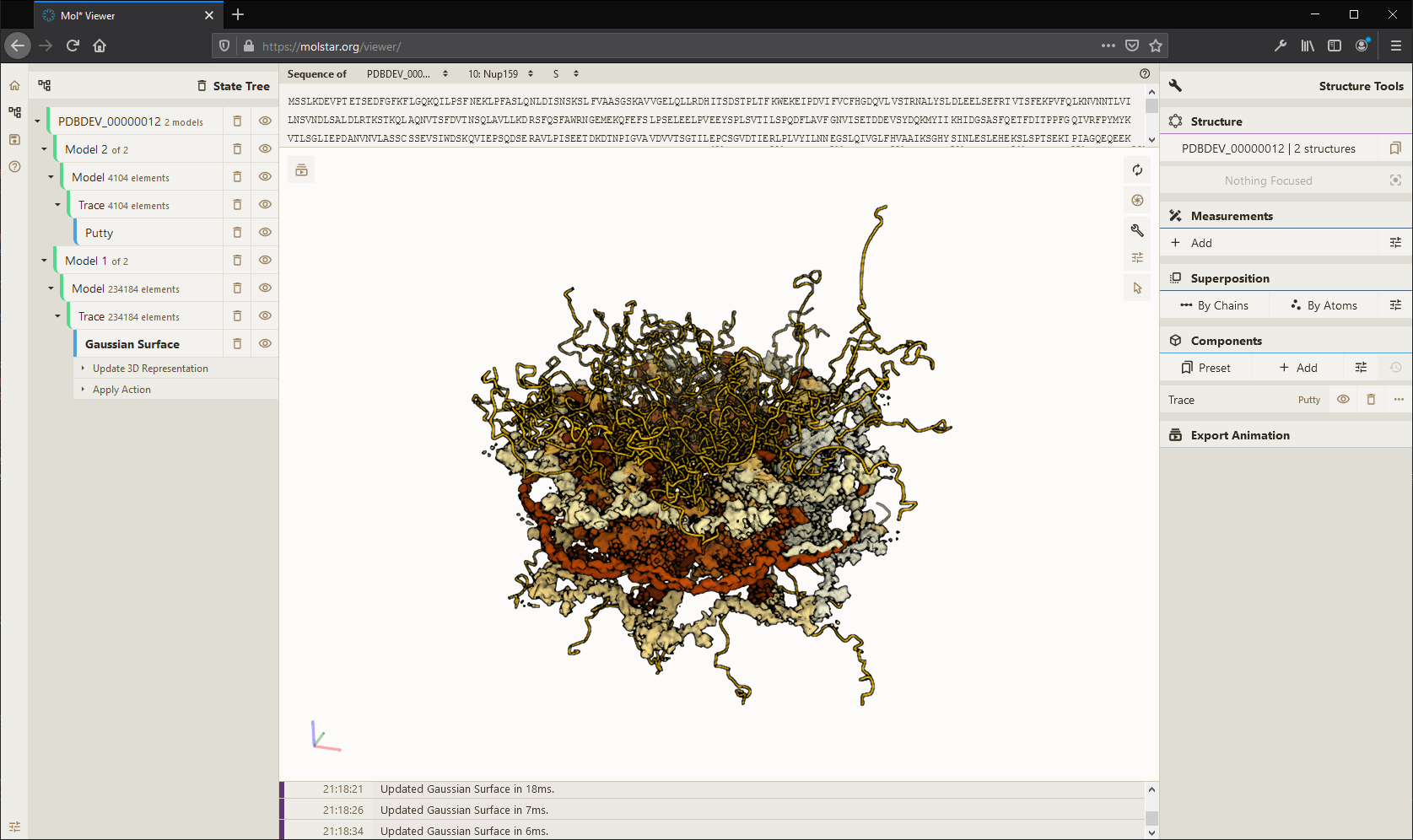
High-performance graphics and data handling of the  Viewer allow users to simultaneously visualize up to hundreds of (superimposed) protein structures, play molecular dynamics trajectories, render cell-level models at atomic detail with tens ofmillions of atoms, or display huge models obtained by I/HM suchas the Nuclear Pore Complex. Read more in NAR.
Viewer allow users to simultaneously visualize up to hundreds of (superimposed) protein structures, play molecular dynamics trajectories, render cell-level models at atomic detail with tens ofmillions of atoms, or display huge models obtained by I/HM suchas the Nuclear Pore Complex. Read more in NAR.
Implemented by: 

Mol*VS
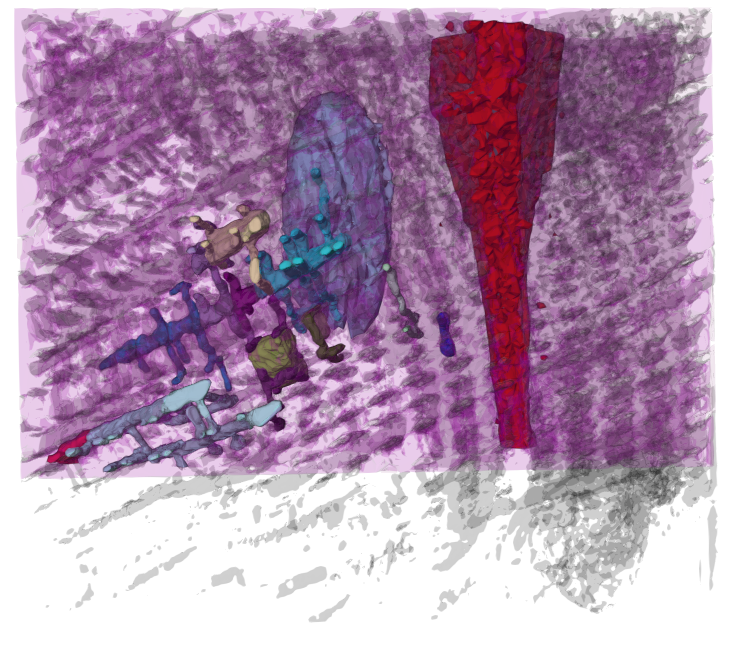
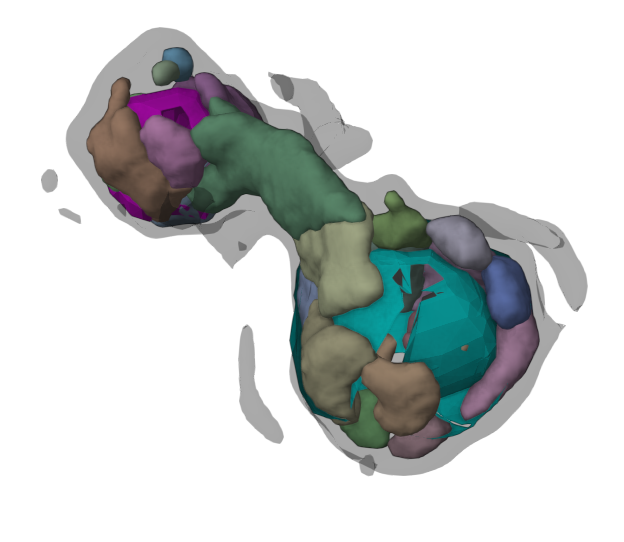
Mol* Volumes & Segmentations (Mol*VS) is a Mol* Viewer extension which adds support for large scale volumetric data & their segmentations. Building on the existing  ecosystem, this extension allows seamless integration of biomolecular data from cellular to atomic scale. Read more in NAR.
ecosystem, this extension allows seamless integration of biomolecular data from cellular to atomic scale. Read more in NAR.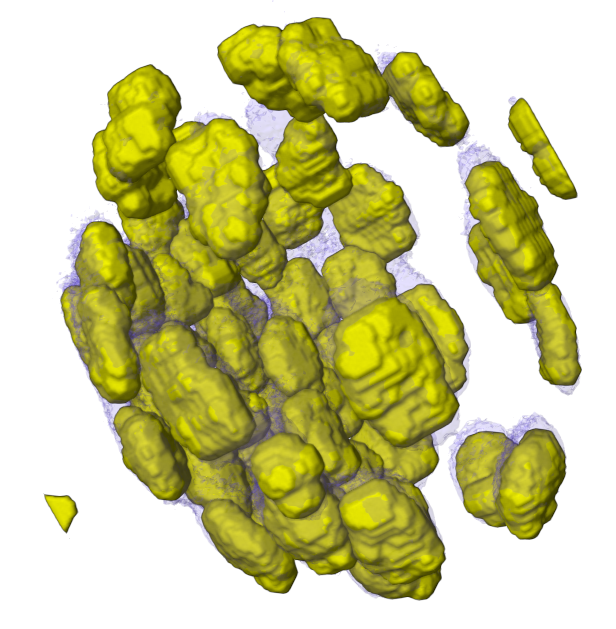

OverProt
 OverProt creates an overview of secondary structure elements in protein families. For each protein family from the CATH database, a secondary structure consensus is available, showing the characteristic helices and β-sheets of the family. Read more in Bioinformatics.
OverProt creates an overview of secondary structure elements in protein families. For each protein family from the CATH database, a secondary structure consensus is available, showing the characteristic helices and β-sheets of the family. Read more in Bioinformatics.
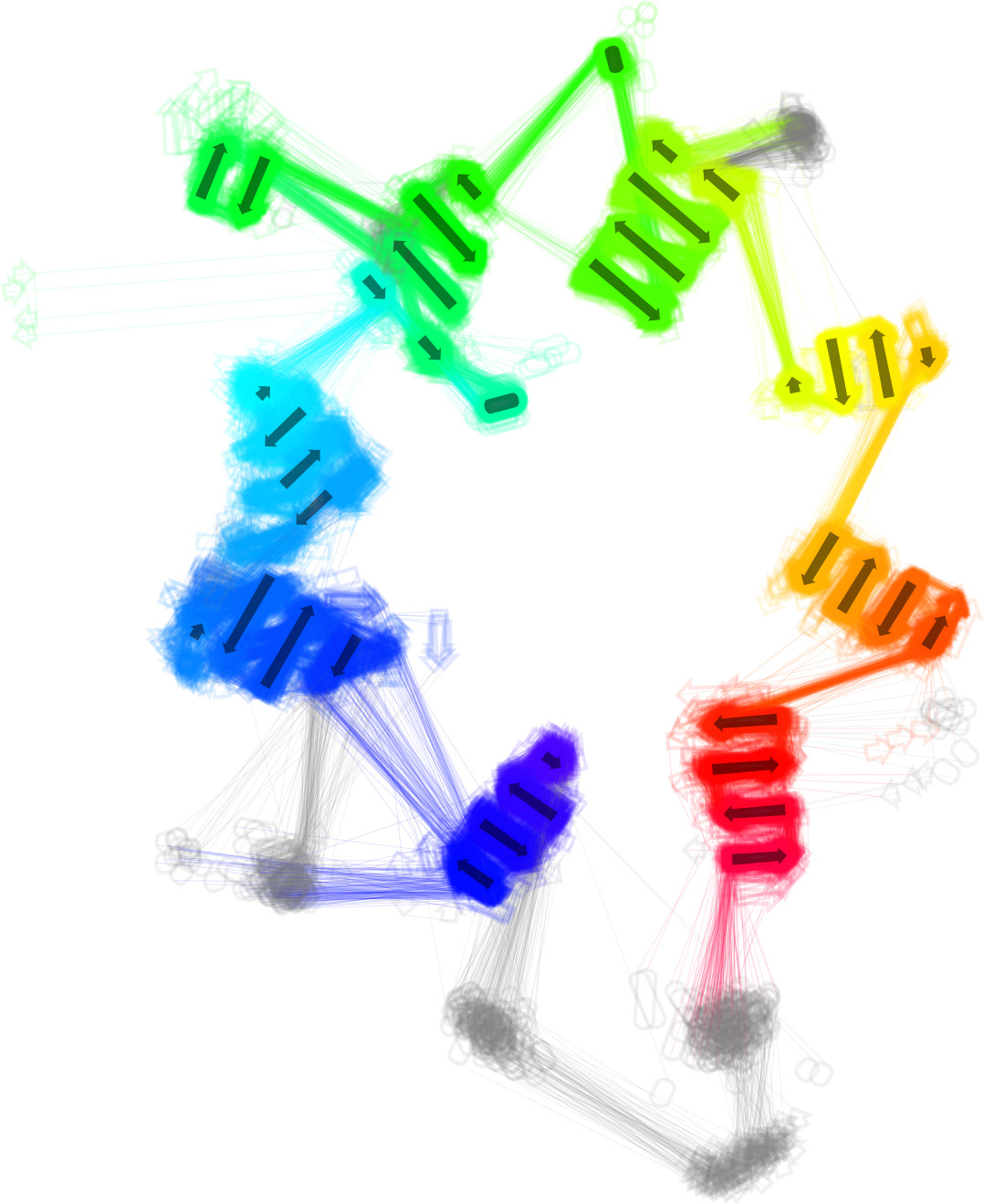
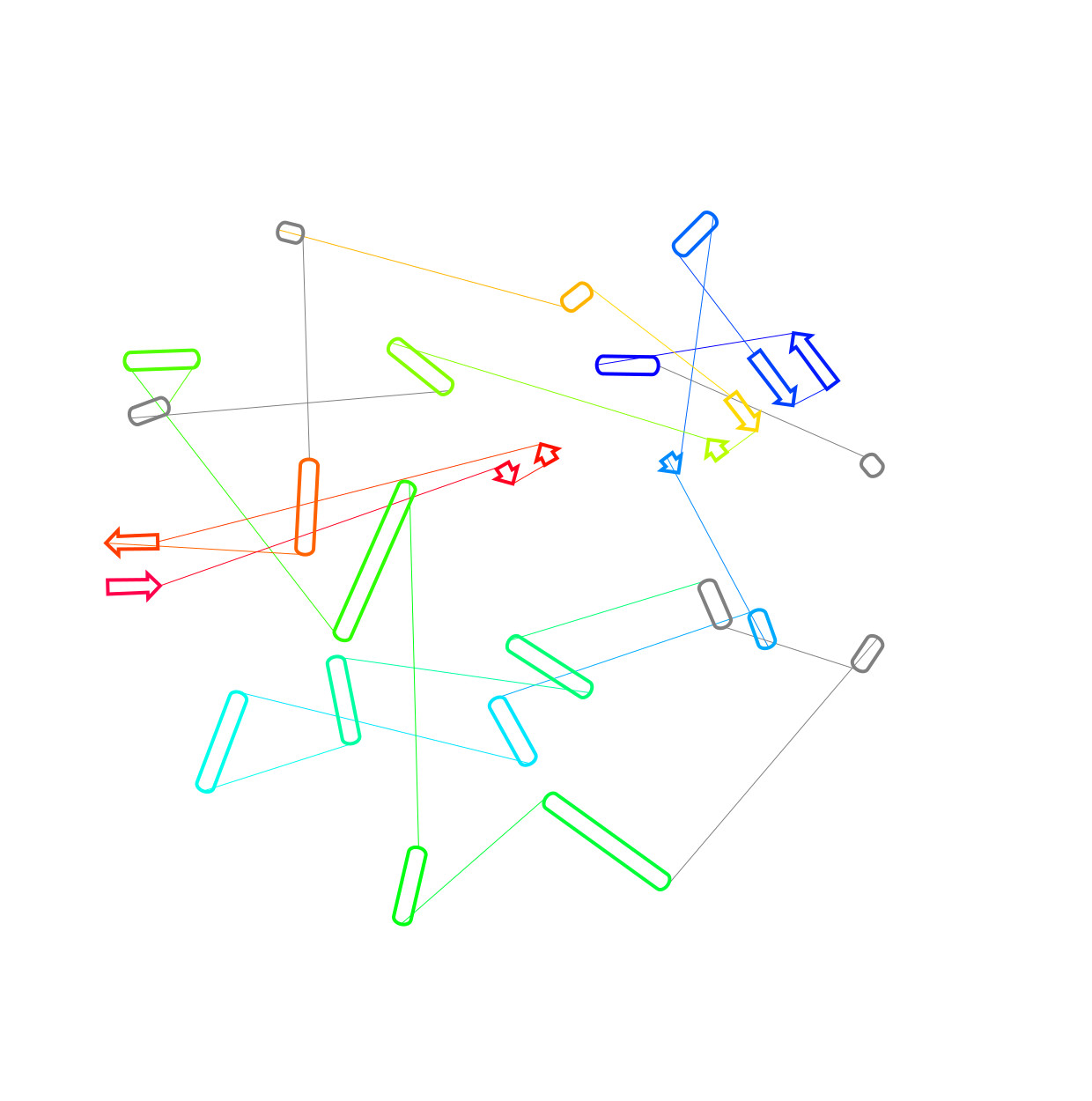
2DProts
2DProts contains 2D diagrams of secondary structures of domains in protein biomacromolecules. Offered diagrams visualize relative distance between single secondary structure elements, as well as position of amino acids that comprise each element in the sequence of the domain. Read more in Bioinformatics.




 biodata(at)ceitec.cz
biodata(at)ceitec.cz